Workflow for High-throughput Screening of Enzyme Mutant Libraries Using Matrix-assisted Laser Desorption/Ionization Mass Spectrometry Analysis of Escherichia coli Colonies
Themes: Conversion
Keywords: Mass Spectrometry
Citation
Choe, K., Sweedler, J.V. Nov. 5, 2023. “Workflow for High-throughput Screening of Enzyme Mutant Libraries Using Matrix-assisted Laser Desorption/Ionization Mass Spectrometry Analysis of Escherichia coli Colonies.” Bio-Protocol. 13(21): e4862. DOI: 10.21769/BioProtoc.4862.
Overview
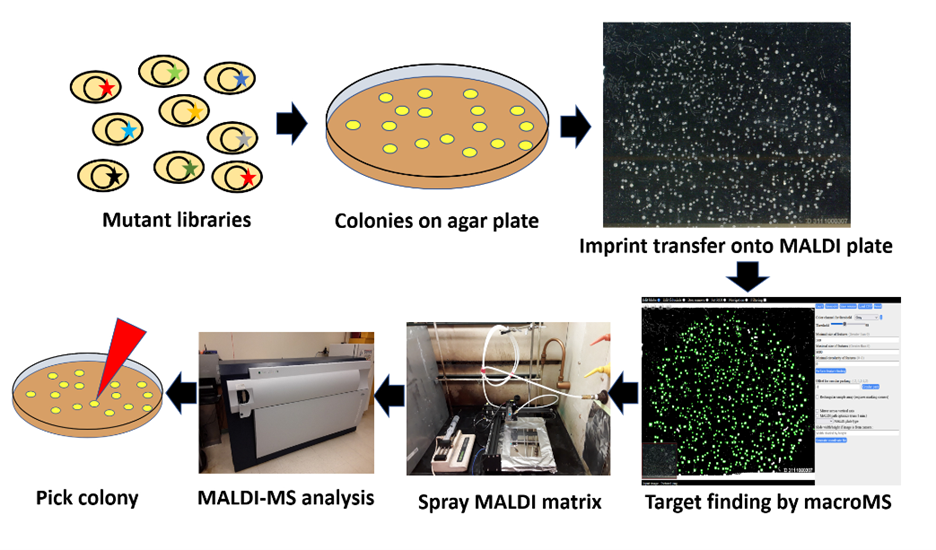
High-throughput molecular screening of microbial colonies and DNA libraries are critical procedures that enable applications such as directed evolution, functional genomics, microbial identification, and creation of engineered microbial strains to produce high-value molecules. A promising chemical screening approach is the measurement of products directly from microbial colonies via optically guided matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS). Measuring the compounds from microbial colonies bypasses liquid culture with a screen that takes approximately 5 s per sample. We describe a protocol combining a dedicated informatics pipeline and sample preparation method that can prepare up to 3,000 colonies in under 3 h. The screening protocol starts from colonies grown on Petri dishes and then transferred onto MALDI plates via imprinting. The target plate with the colonies is imaged by a flatbed scanner and the colonies are located via custom software. The target plate is coated with MALDI matrix, MALDI-MS analyzes the colony locations, and data analysis enables the determination of colonies with the desired biochemical properties. This workflow screens thousands of colonies per day without requiring additional automation. The wide chemical coverage and the high sensitivity of MALDI-MS enable diverse screening projects such as modifying enzymes and functional genomics surveys of gene activation/inhibition libraries.