KETCHUP: Parameterizing of Large-Scale Kinetic Models Using Multiple Datasets with Different Reference States
Themes: Conversion
Keywords: Metabolomics, Modeling
Citation
Hu, M., Suthers, P.F., Maranas, C.D. Feb. 7, 2024. “Kinetic Estimation Tool Capturing Heterogeneous Datasets Using Pyomo (KETCHUP)” GitHub Repository.
Overview
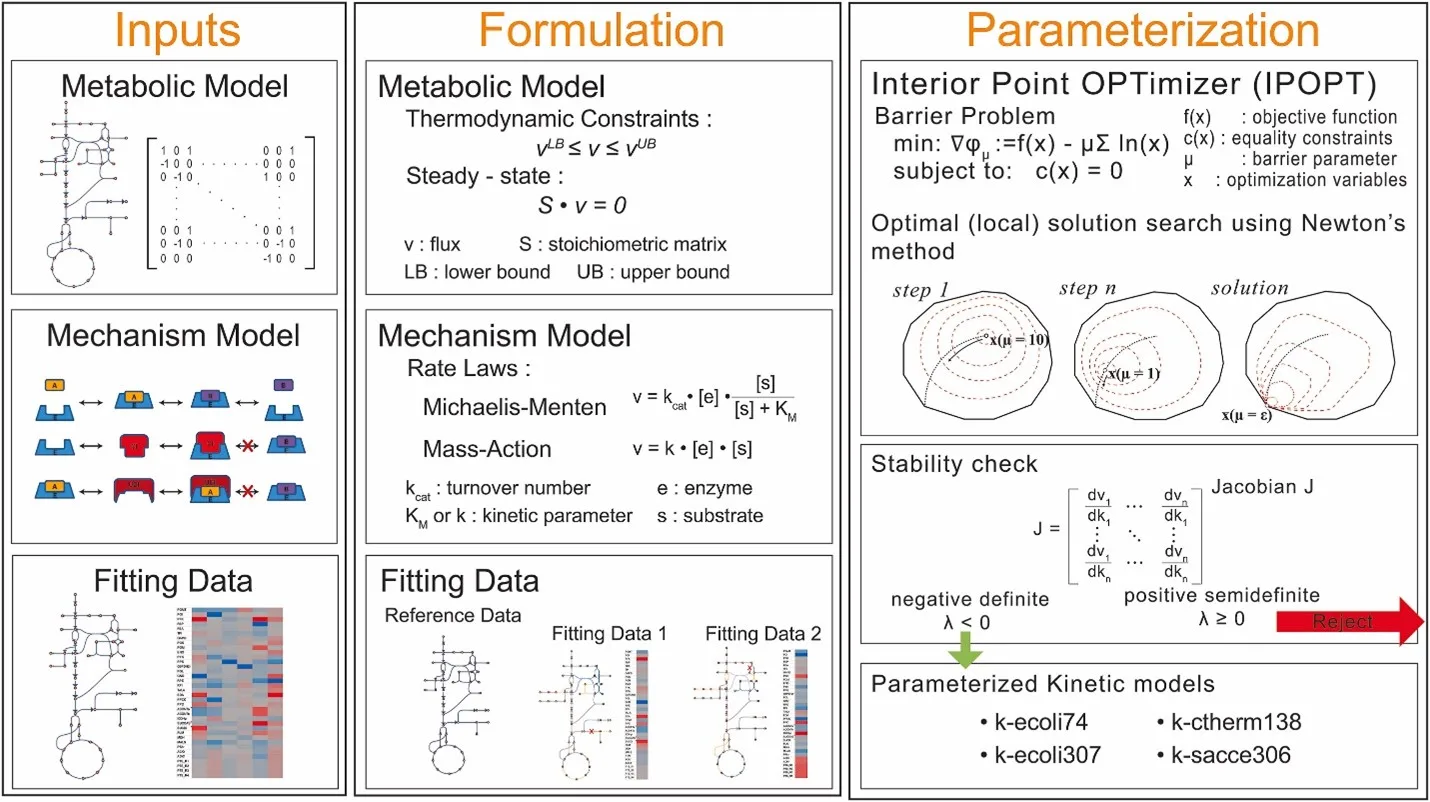
Repository for Kinetic Estimation Tool Capturing Heterogeneous Datasets Using Pyomo (KETCHUP), a flexible parameter estimation tool that leverages a primal-dual interior-point algorithm to solve a nonlinear programming (NLP) problem that identifies a set of parameters capable of recapitulating the steady-state fluxes and concentrations in wild-type and perturbed metabolic networks.
KETCHUP can use K-FIT [2] input files. Example K-FIT input files are located in the K-FIT repository at https://github.com/maranasgroup/K-FIT.
Data
GitHub Repository: Includes code and updated installation guide.
Illinois Data Bank includes:
- Model statistics
- Strain metabolites and fluxomics data
- Yield prediction comparisons