Data for A Generalized Platform for Artificial Intelligence-powered Autonomous Protein Engineering
Themes: Conversion
Keywords: AI/ML, Automation
Citation
Singh, N., Lane, S., Yu, T., Lu, J., Ramos, A., Cui, H., Zhao, H. July 1, 2025. Data for: “A Generalized Platform for Artificial Intelligence-powered Autonomous Enzyme Engineering.” Zenodo. DOI: 10.5281/zenodo.15243670.
Overview
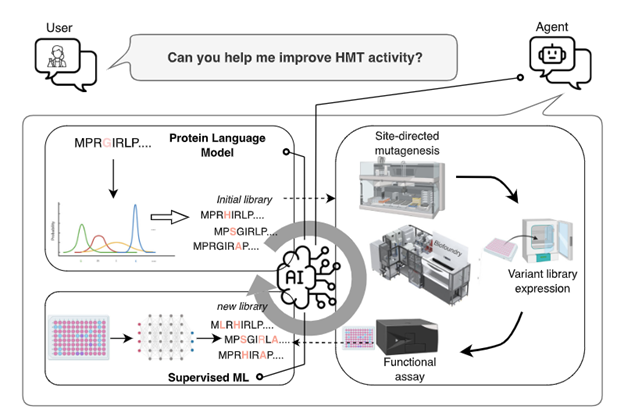
This repository accompanies the work “A Generalized Platform for Artificial Intelligence-powered Autonomous Enzyme Engineering”.
Proteins are the molecular machines of life with numerous applications in energy, health, and sustainability. However, engineering proteins with desired functions for practical applications remains slow, expensive, and specialist-dependent. Here we report a generally applicable platform for autonomous enzyme engineering that integrates machine learning and large language models with biofoundry automation to eliminate the need for human intervention, judgement, and domain expertise. Requiring only an input protein sequence and a quantifiable way to measure fitness, this automated platform can be applied to engineer a wide array of proteins. As a proof of concept, we engineer Arabidopsis thaliana halide methyltransferase (AtHMT) for a 90-foldimprovement in substrate preference and 16-fold improvement in ethyl-transferase activity, along with developing a Yersinia mollaretii phytase (YmPhytase) variant with 26-fold improvement in activity at neutral pH. This is accomplished in four rounds over 4 weeks, while requiring construction and characterization of fewer than 500 variants for each enzyme. This platform for autonomous experimentation paves the way for rapid advancements across diverse industries, from medicine and biotechnology to renewable energy and sustainable chemistry.
Data
Zenodo: Primer design and worklists
Zenodo: Python scripts
Illinois Data Bank: Raw data and experiment video